Introduction
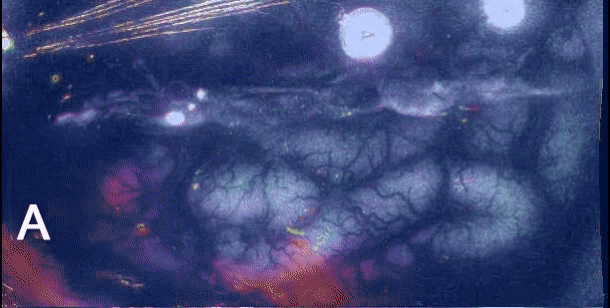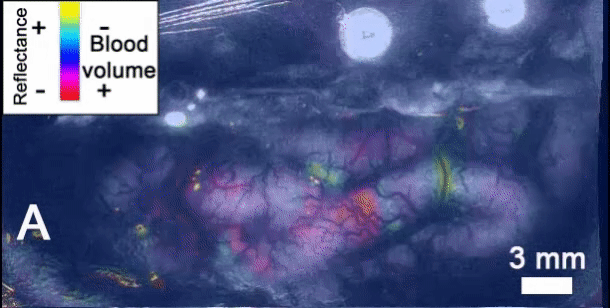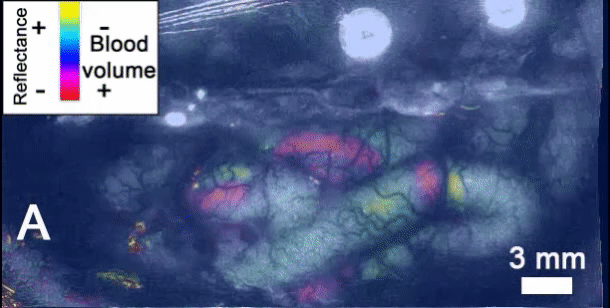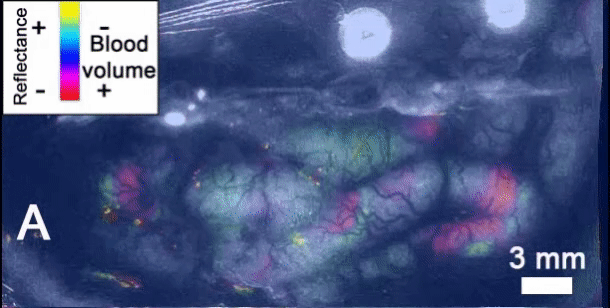
Type 3 familial hemiplegic migraine (FHM3) is an autosomal dominant disease characterized by migraine with aura that causes motor impairment on one side of the body (hemiplegia) in addition to an aura before the migraine begins. FHM3 is one of four subtypes, caused by mutations in the gene SCN1A [1]. Auras and hemiplegia from FHM3 are believed to be caused by cortical spreading depression (CSD). This is a wave of neuronal depolarization that slowly propagates through the brain [1]. Part of this is an influx of sodium into neurons, which causes water to enter cells. This causes cells to swell and extracellular space to be reduced [2].
FHM3 and SCN1A
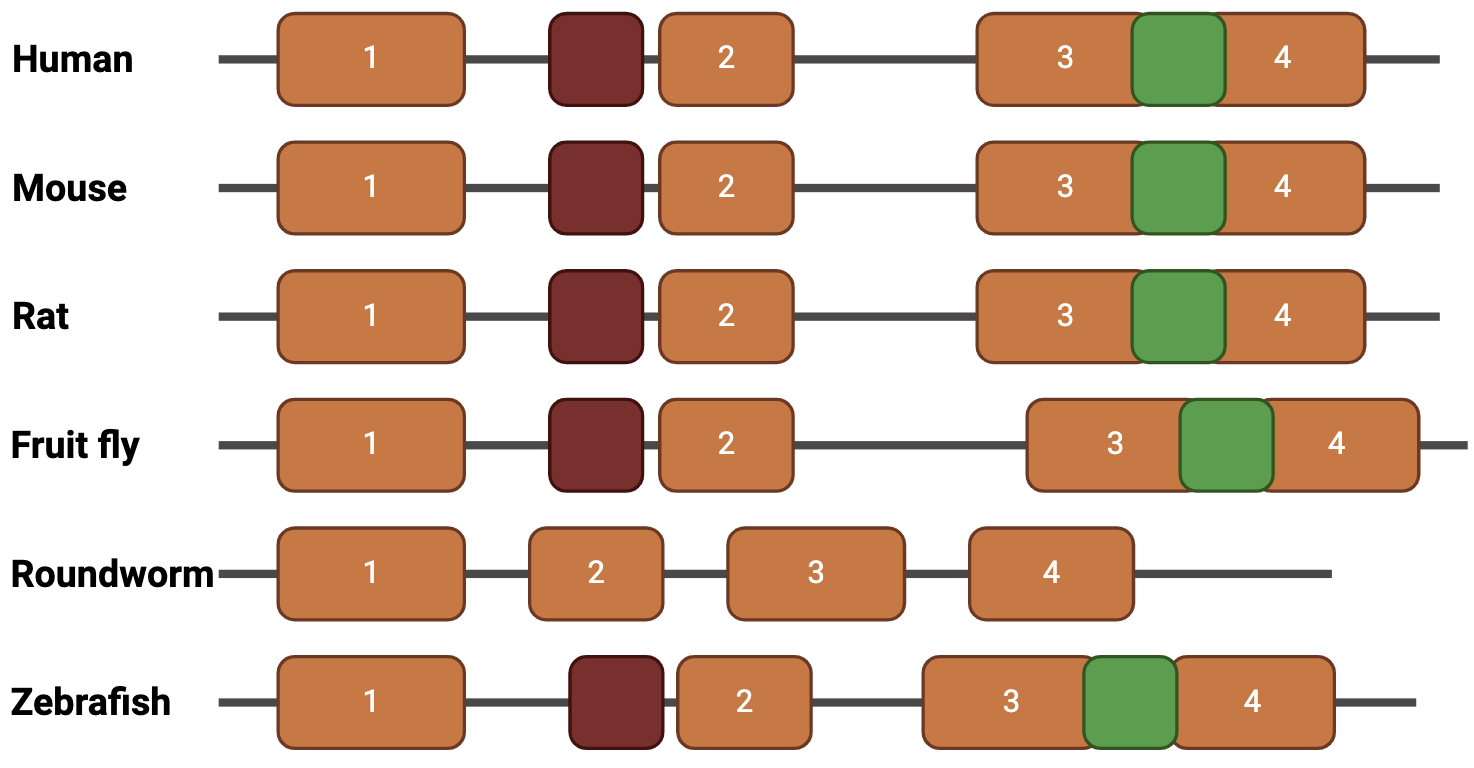
FHM3 is characterized by gain-of-function mutations in the gene SCN1A, which is located on chromosome 2 [3]. SCN1A consists of multiple domains. These include ion transport domains, a cytoplasmic domain, and a linker between domains 3 and 4. This linker region is responsible for the inactivation of the voltage-gated sodium channel, Nav1.1, coded by SCN1A [4].
Gene Ontology for SCN1A
There are quite a few gene ontology terms associated with SCN1A, most relating to action potential propagation and the voltage-gated sodium channel complex. These are useful for this project because the believed cause behind migraine auras, CSD, is a wave of depolarization and repolarization propagating through the brain. This can be caused by defective sodium channels allowing too much sodium into neurons.
|
Molecular function [5]
|
Biological process [5]
|
Cellular component [5]
|
Gap in knowledge
While SCN1A has been studied extensively for Dravet's syndrome [6], it has not been studied much for FHM3 due to its rarity. Because of this, its role in causing hemiplegia is not fully understood.
Mice are a great model organism for studying FHM3
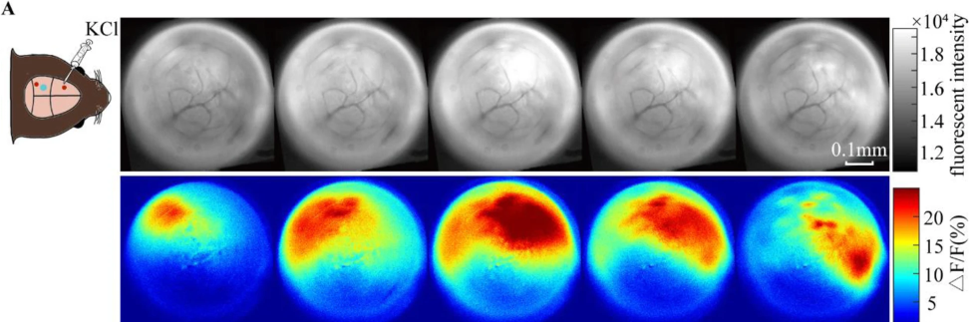
Mus musculus (mice) will be used to study FHM3. CSD can easily be induced in mice through topical application of KCl [7]. This will help me to ensure the mice experience hemiplegia. Additionally, behavioral assays can be done through a variety of ways, but for the purposes of this experiment, I have chosen the open field test to examine motor impairment in mice. These tests are typically used to measure locomotion and anxiety [8]. Mice are placed in an open white plexiglass box, then movement is tracked through an automated tracking system [9]. Mice that are experiencing hemiplegia will only be able to walk in small circles, due to half of their body being paralyzed. Wild type mice should have normal walking patterns like the one shown on the left in Figure ().
SCN1A Protein Interactions
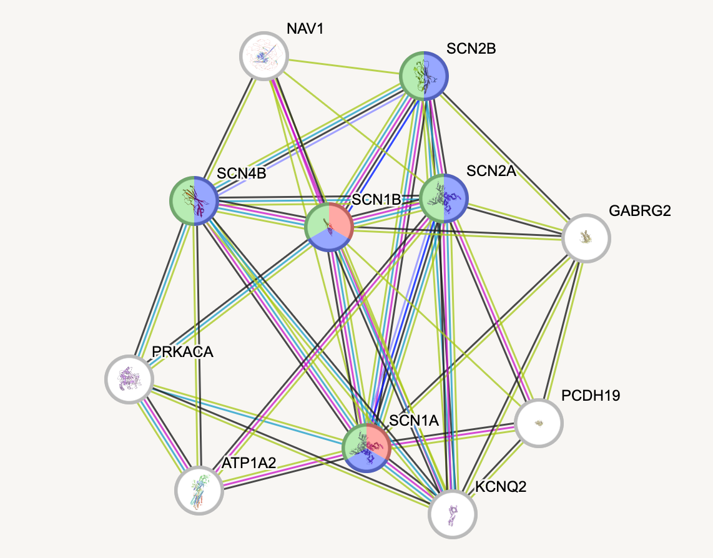
The protein interaction networks between human and mouse SCN1A are quite similar. As you can see in the figure below, they are involved in the same biological processes, molecular functions, and cellular components. While the mice have a few more protein interactions, their networks includes all the genes highlighted in the human one. Protein interaction networks were obtained from the String database.
Biological process: action potential propagation
Molecular functions: voltage-gated sodium channel activity
Cellular components: plasma membrane
Biological process: action potential propagation
Molecular functions: voltage-gated sodium channel activity
Cellular components: plasma membrane
Primary Goal
My primary goal is to investigate how mutations in SCN1A cause fully reversible motor impairment. The linker region between domains 3 and 4 is of particular interest in the case because it is responsible for the inactivation of the gate. I will complete this goal through the following aims:
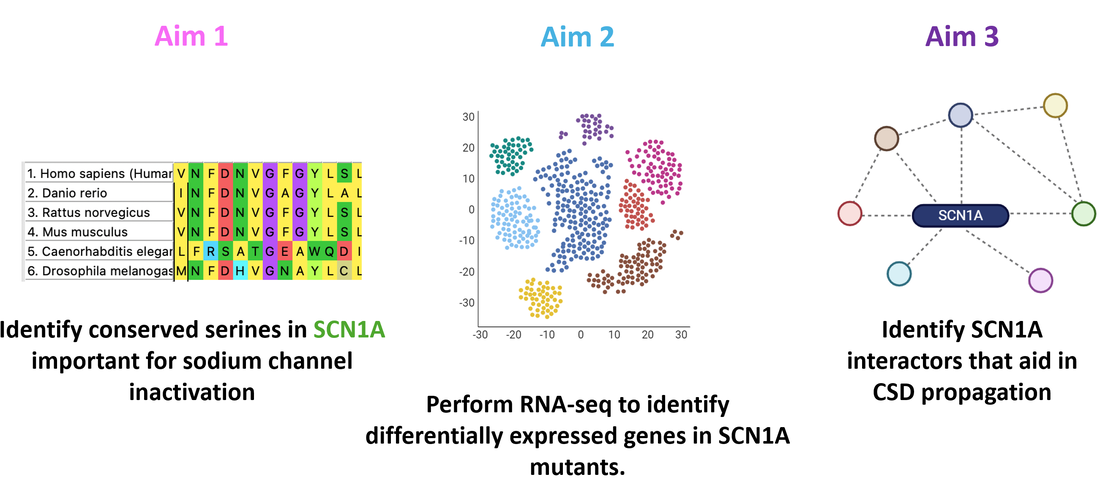
Aim 1
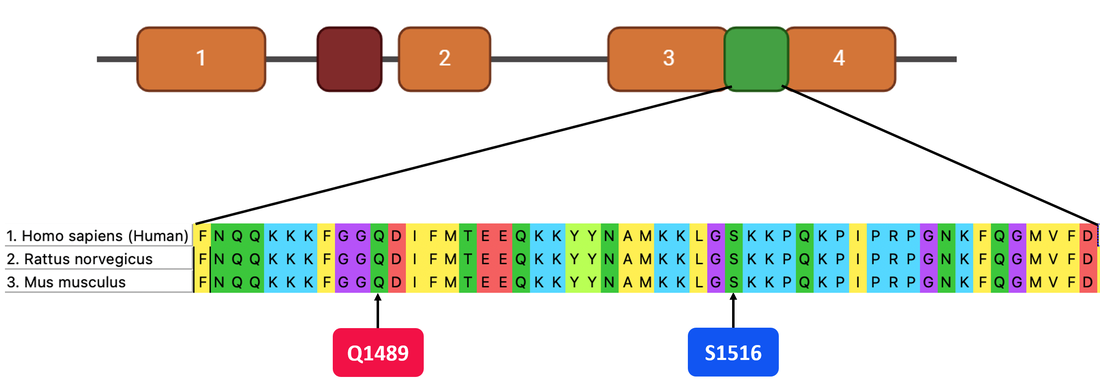
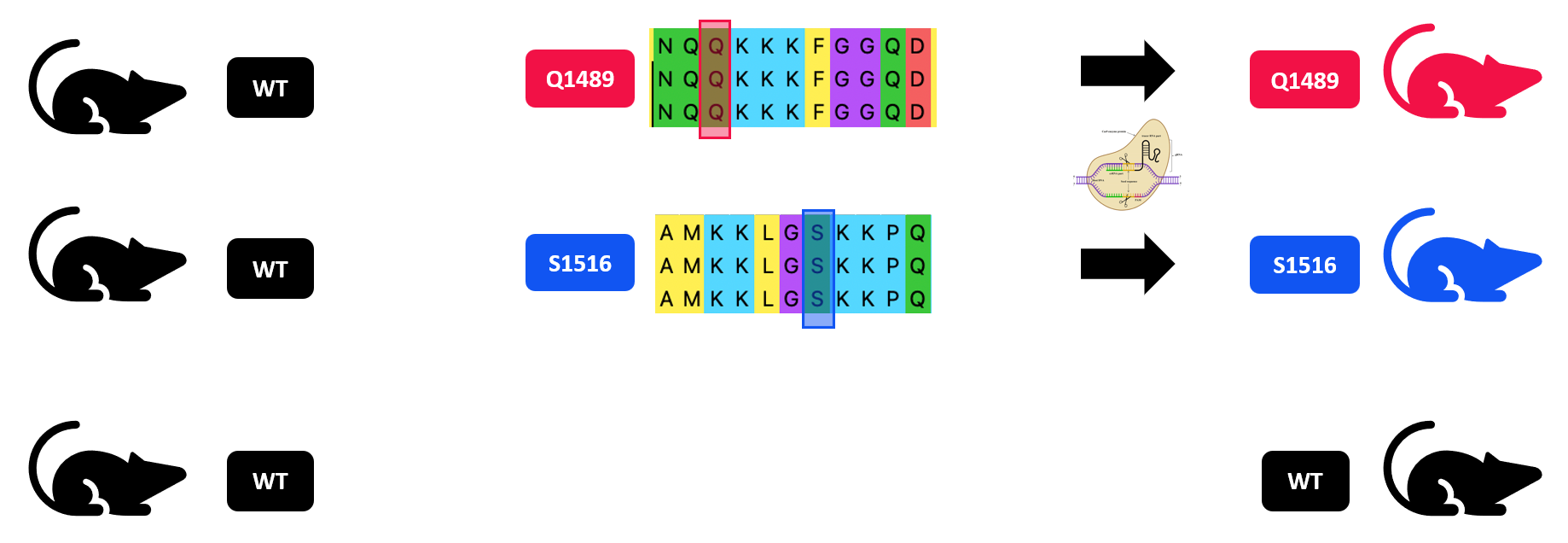
In aim 1, I will perform a multiple sequence alignment using FASTA data obtained through Ensembl, MEGA, and Clustal Omega. I will then choose different conserved points in this region to mutate. Q1489K is a known mutation that causes FHM3, so I have picked that as a control. In addition to that site, I will mutate S1516. Using UniProt, I found that phosphorylation at this serine is involved in slowing inactivation of this gate, as well as reducing peak sodium currents [10]. This leads me to believe that mutating it may have some affect on CSD induction.
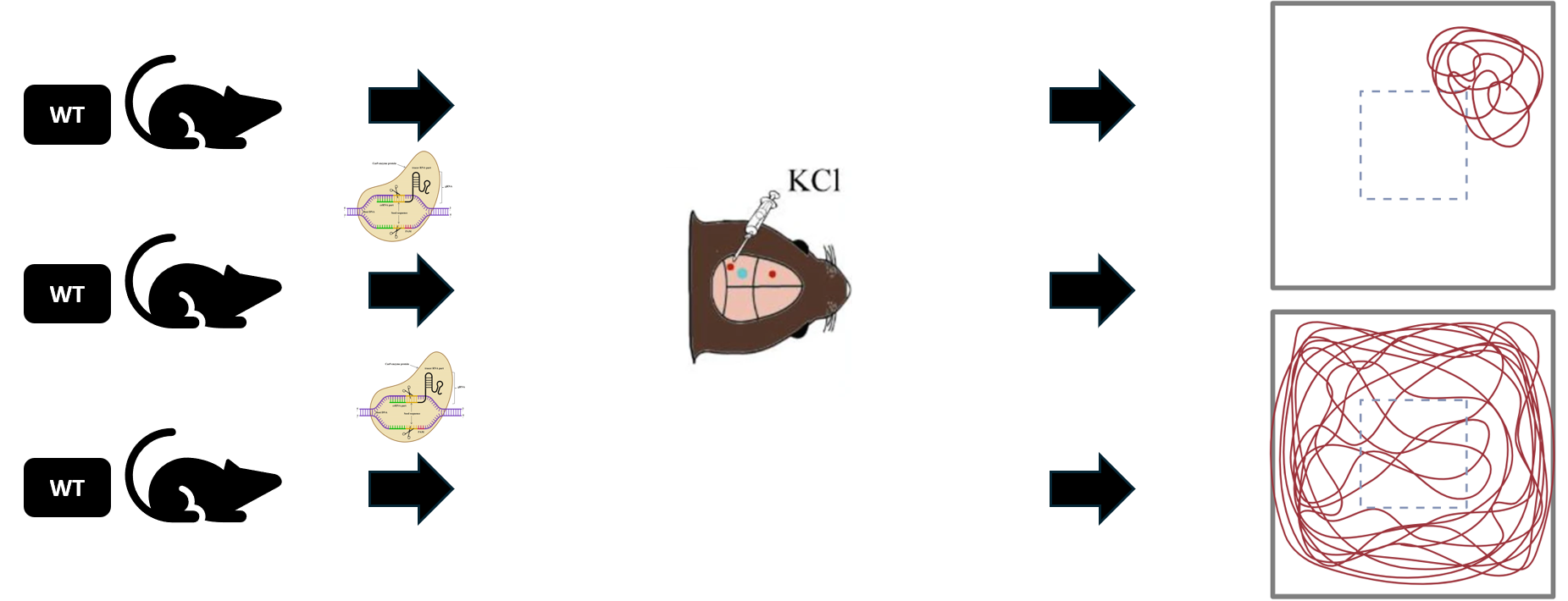
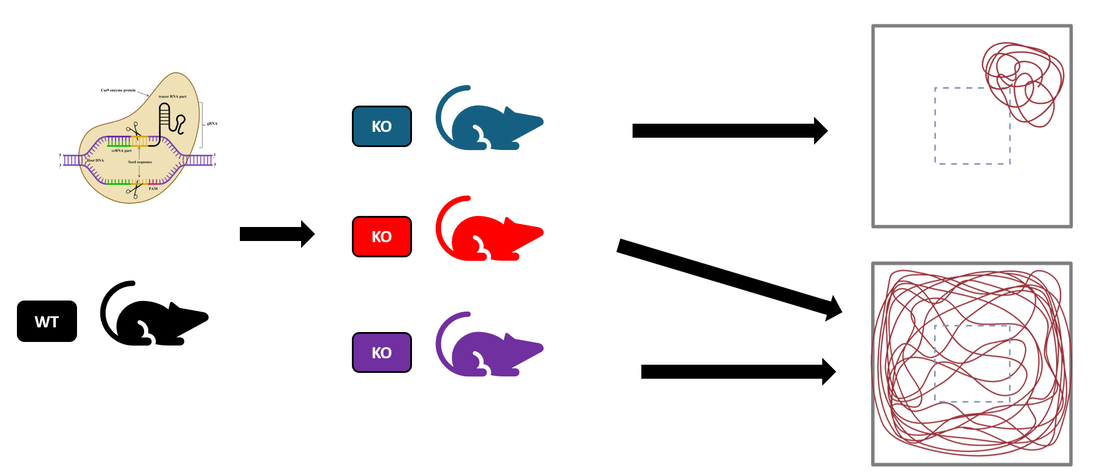
I will use CRISPR/Cas9 to knock in mutations at these conserved points. I will also leave wild type (WT) mice as a control group.
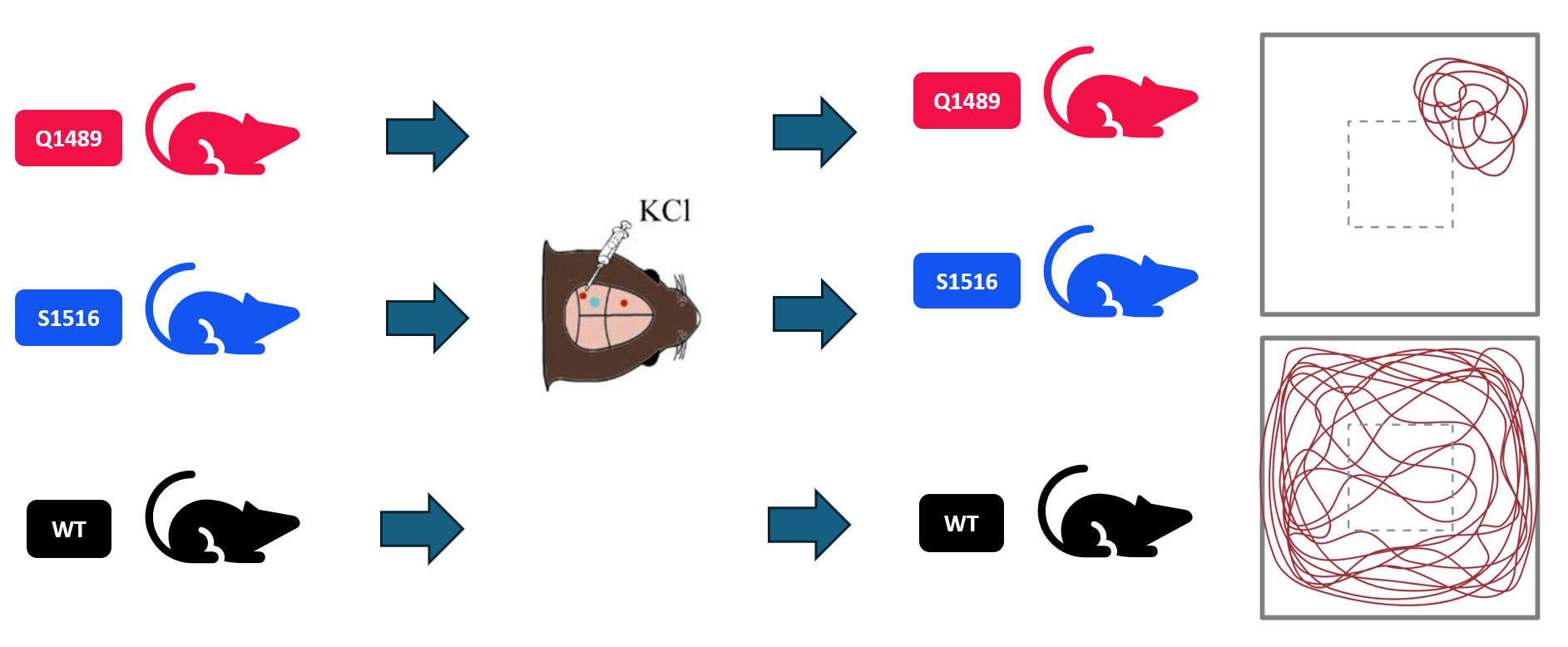
Finally, I will induce CSD in these three groups of mice and run the open field test to assess motor impairment. I hypothesize that once these mutations are introduced to the mice, they will have a lower threshold needed to induce CSD. They should need less KCl to induce CSD than the WT mice need. The same amount of KCl will be applied to both WT and mutant mice until the mutants are experiencing CSD, at which point open field tests will be performed to analyze for hemiplegia. The mutant mice should be experiencing hemiplegia, and will have a walking pattern like the one shown on top below. The WT mice should not be experiencing CSD, therefore no hemiplegia, and should have a normal pattern such as the one on the bottom.
Aim 2
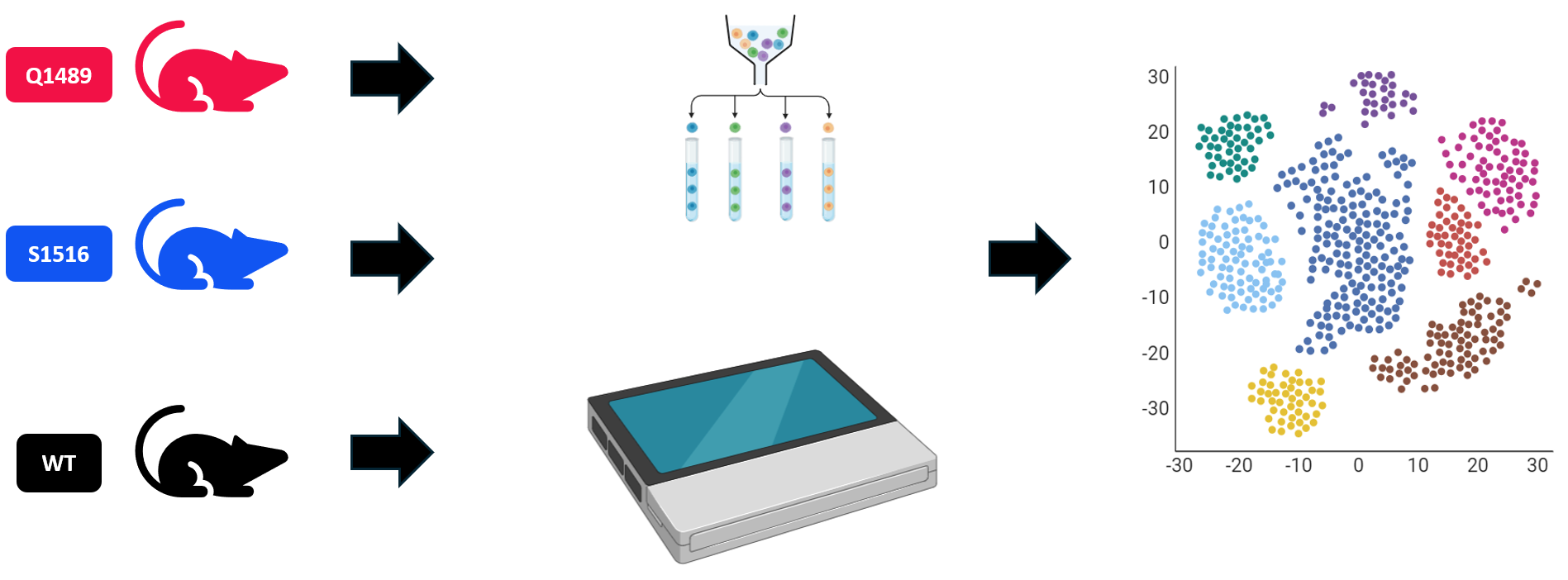
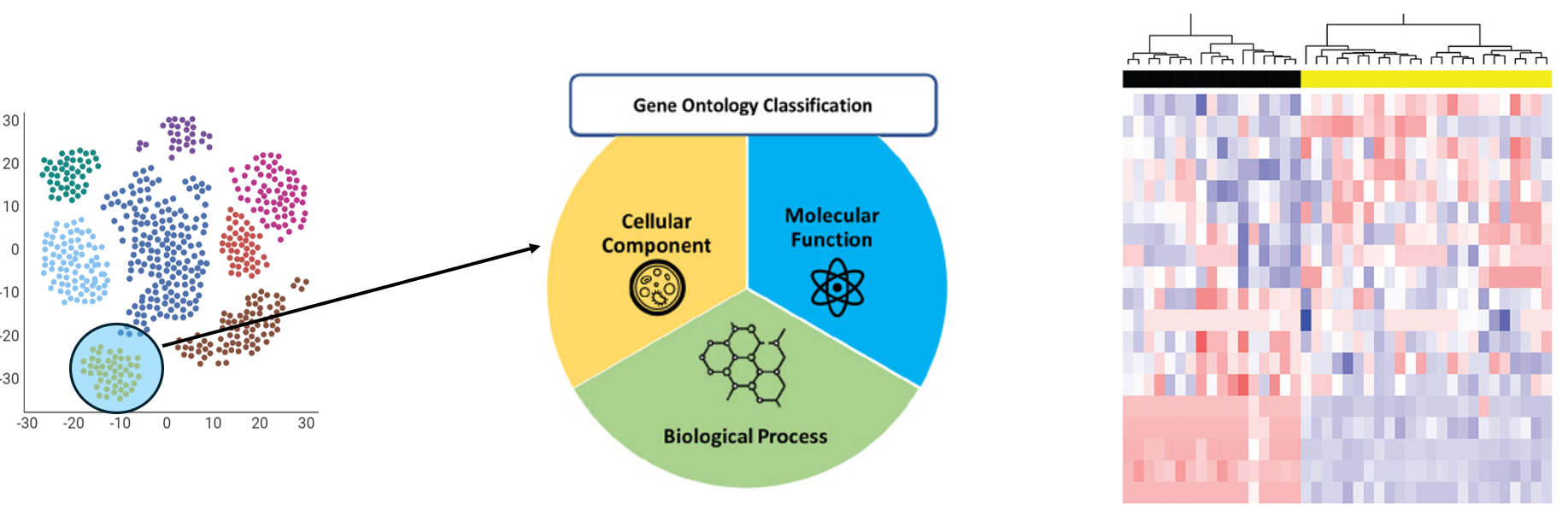
In aim 2, I will perform single cell RNA-sequencing (scRNAseq) to identify differential gene expression in controls compared to mutants. I will obtain tissue from the brains of mutants and controls and use KCl to induce CSD in all samples. From there, I will perform scRNAseq to get a Principle Component Analysis (PCA).
I will analyze the PCA using Gene Ontology terms, then use a heat map to see which genes are upregulated and downregulated in mutants compared to controls. I will be specifically looking for genes that are downregulated whose biological processes also include action potential propagation.
Finally, I will validate results from this aim using CRISPR/Cas9 and open field tests once again. Using WT mice, I will knock out genes that were downregulated in mutant mice. Following this, I will induce CSD using the amount of KCl it took to induce it in mutant mice. I hypothesize that some of the genes that were downregulated in mutant mice will cause mutant-like phenotyping (hemiplegia) when KCl is applied.
Aim 3
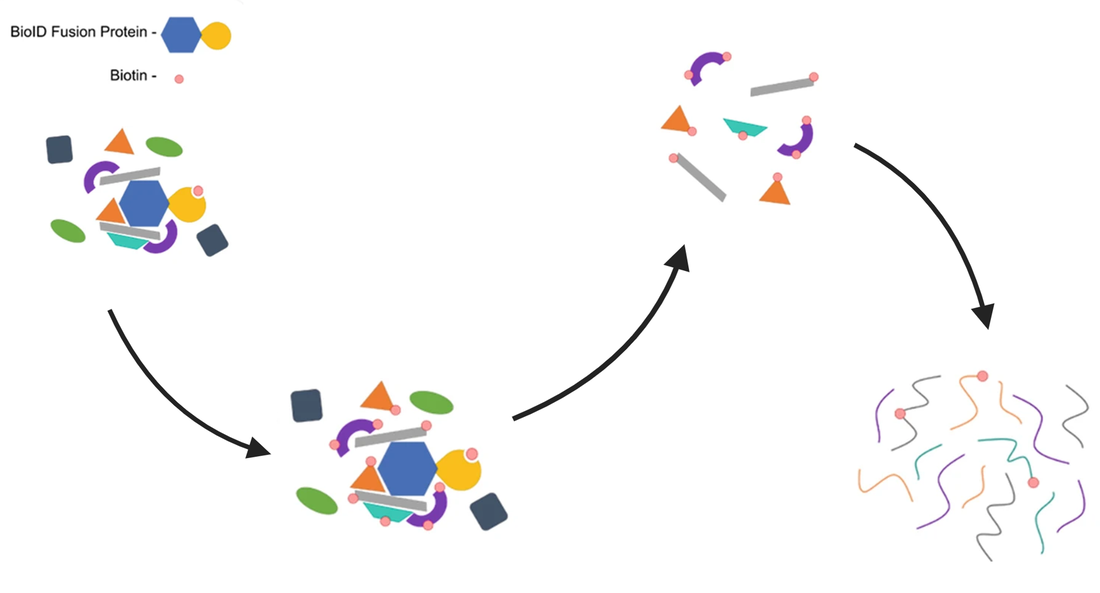
In aim 3, I will identify SCN1A interactor proteins that aid in CSD propagation, which can then lead to hemiplegia. First, I will use BioID to identify proteins that SCN1A interacts with in the brain. Proteins that interact with SCN1A will be biotinylated and then isolated through the use of streptavidin beads. This is shown in the Figure ().
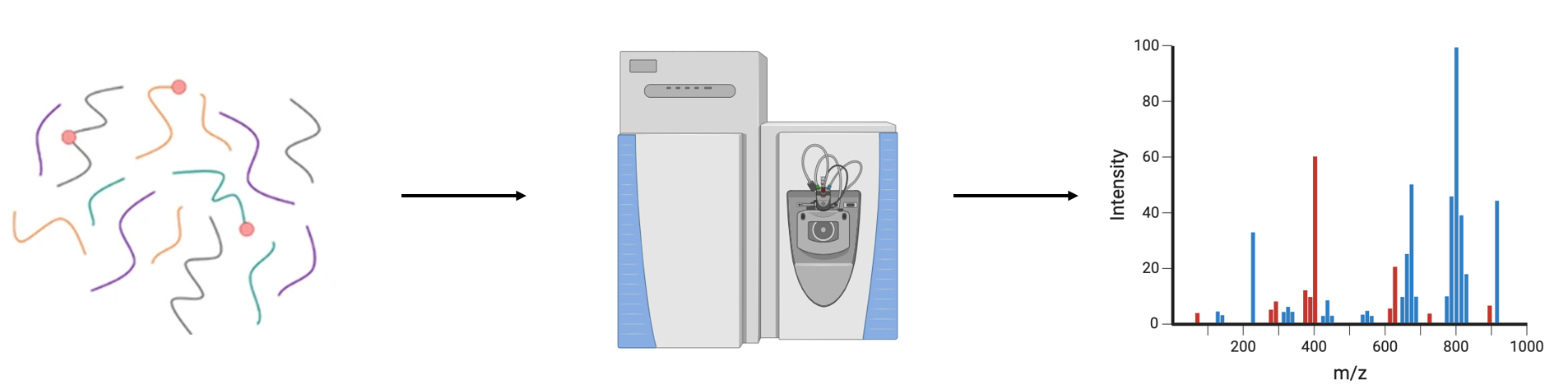
I will then take the isolated proteins and perform mass spectrometry on them to identify them. I am looking for proteins that are downregulated in mutant mice from Aim 1 compared to the controls.
Finally, I will validate the results from this aim using CRISPR/Cas9 for one final time. I will take WT mice and knockout proteins that were not found or downregulated in the mutant mice from Aim 1. I hypothesize that most of the proteins knocked out will not have an effect, and these mice will still have the WT pattern in the open field test. However, I believe there may be some proteins that when knocked out, leave the mice with mutant-like phenotyping, who do have hemiplegia and can only walk in small circles when CSD is induced.
Conclusions
Type III Familial Hemiplegic Migraine is caused by gain of function mutations in the SCN1A gene, and symptoms include hemiplegia (fully reversible motor impairment). Due to rarity of this disease, the role of SCN1A in motor function is not fully known. Using genomic and proteomic techniques as I have demonstrated in my Aims above can help us to fill in this gap in knowledge and also may provide insight into therapeutic targets for symptoms of FHM3 such as hemiplegia.
Future Directions
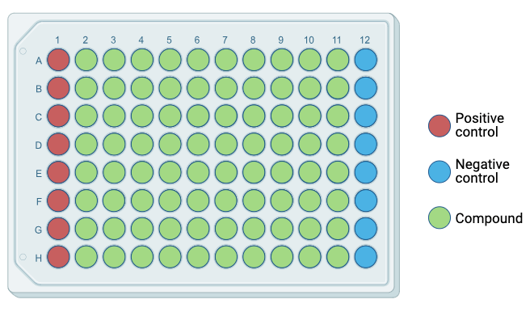
My proposal for future directions involves and forward chemical screen using a targeted library based off of a compound called GS967. This compound is a sodium channel inhibitor that has previously been found to bind to Nav1.1, the channel that SCN1A encodes [11]. I would like to make a targeted library of compounds similar to GS967 to assess for phenotypic rescue, with trials beginning in zebrafish larvae. I believe this compound, or others like it, show promise to be an effective therapy for those that get FHM3.
Drafts
| final_presentation.pdf | |
| File Size: | 3147 kb |
| File Type: | |
References
[1] Kazemi, H., Speckmann, E.-J., & Gorji, A. (2014). Familial Hemiplegic Migraine and Spreading Depression. Iranian Journal of Child Neurology, 8(3). https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4135274/
[2] James, M. F., Smith, J. M., Boniface, S. J., Huang, C. L.-H., & Leslie, R. A. (2001). Cortical spreading depression and migraine: New insights from imaging? Trends in Neurosciences, 24(5), 266–271. https://doi.org/10.1016/s0166-2236(00)01793-8
[3] https://www.ncbi.nlm.nih.gov/gene/6323
[4] https://www.ebi.ac.uk/interpro/entry/cdd/CD13433/
[5]InterProScan Search Result, SCN1A. InterPro (n.d.). https://www.ebi.ac.uk/interpro/result/InterProScan/iprscan5-R20240403-224305-0683-43191542-p1m/
[6] Bender, A. C., Morse, R. P., Scott, R. C., Holmes, G. L., & Lenck-Santini, P.-P. (2012). SCN1A mutations in Dravet Syndrome: Impact of Interneuron dysfunction on neural networks and cognitive outcome. Epilepsy & Behavior, 23(3), 177–186. https://doi.org/10.1016/j.yebeh.2011.11.022
[7] Fu, X., Chen, M., Lu, J., & Li, P. (2022). Cortical spreading depression induces propagating activation of the thalamus ventral posteromedial nucleus in awake mice. The Journal of Headache and Pain, 23(1). https://doi.org/10.1186/s10194-021-01370-z
[8] Seibenhener, M. L., & Wooten, M. C. (2015). Use of the open field maze to measure locomotor and anxiety-like behavior in mice. Journal of Visualized Experiments, (96). https://doi.org/10.3791/52434
[9] Stanford Med. Sensorimotor tests, Open Field. https://med.stanford.edu/sbfnl/services/bm/sm/openfield.html
[10] UniProt search results, SCN1A. https://www.uniprot.org/uniprotkb/P35498/entry
[11] Barbieri, R., Bertelli, S., Pusch, M., & Gavazzo, P. (2019). Late sodium current blocker GS967 inhibits persistent currents induced by familial hemiplegic migraine type 3 mutations of the SCN1A gene. The Journal of Headache and Pain, 20(1). https://doi.org/10.1186/s10194-019-1056-2
[2] James, M. F., Smith, J. M., Boniface, S. J., Huang, C. L.-H., & Leslie, R. A. (2001). Cortical spreading depression and migraine: New insights from imaging? Trends in Neurosciences, 24(5), 266–271. https://doi.org/10.1016/s0166-2236(00)01793-8
[3] https://www.ncbi.nlm.nih.gov/gene/6323
[4] https://www.ebi.ac.uk/interpro/entry/cdd/CD13433/
[5]InterProScan Search Result, SCN1A. InterPro (n.d.). https://www.ebi.ac.uk/interpro/result/InterProScan/iprscan5-R20240403-224305-0683-43191542-p1m/
[6] Bender, A. C., Morse, R. P., Scott, R. C., Holmes, G. L., & Lenck-Santini, P.-P. (2012). SCN1A mutations in Dravet Syndrome: Impact of Interneuron dysfunction on neural networks and cognitive outcome. Epilepsy & Behavior, 23(3), 177–186. https://doi.org/10.1016/j.yebeh.2011.11.022
[7] Fu, X., Chen, M., Lu, J., & Li, P. (2022). Cortical spreading depression induces propagating activation of the thalamus ventral posteromedial nucleus in awake mice. The Journal of Headache and Pain, 23(1). https://doi.org/10.1186/s10194-021-01370-z
[8] Seibenhener, M. L., & Wooten, M. C. (2015). Use of the open field maze to measure locomotor and anxiety-like behavior in mice. Journal of Visualized Experiments, (96). https://doi.org/10.3791/52434
[9] Stanford Med. Sensorimotor tests, Open Field. https://med.stanford.edu/sbfnl/services/bm/sm/openfield.html
[10] UniProt search results, SCN1A. https://www.uniprot.org/uniprotkb/P35498/entry
[11] Barbieri, R., Bertelli, S., Pusch, M., & Gavazzo, P. (2019). Late sodium current blocker GS967 inhibits persistent currents induced by familial hemiplegic migraine type 3 mutations of the SCN1A gene. The Journal of Headache and Pain, 20(1). https://doi.org/10.1186/s10194-019-1056-2
About the website
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Joely Swanson, [email protected]
Last updated April 18th, 2024
Genetics 564 website
Joely Swanson, [email protected]
Last updated April 18th, 2024
Genetics 564 website