What is phylogeny?
Phylogenetic trees are a visual representation of the evolutionary relationships in a group of organisms. These are useful as they show how different species are related to each other. They are based on shared ancestry and show how closely related homologs are to one another.
These trees can be made using a combination of different softwares and websites. After obtaining FASTA sequences of orthologues from Ensembl, you can perform a multiple sequence alignment using MEGA. After this, you can create a tree using the bootstrapping option, which shows you how many times out of 100 the same branches resulted when creating a phylogenetic construction.
These trees can be made using a combination of different softwares and websites. After obtaining FASTA sequences of orthologues from Ensembl, you can perform a multiple sequence alignment using MEGA. After this, you can create a tree using the bootstrapping option, which shows you how many times out of 100 the same branches resulted when creating a phylogenetic construction.
Discussion
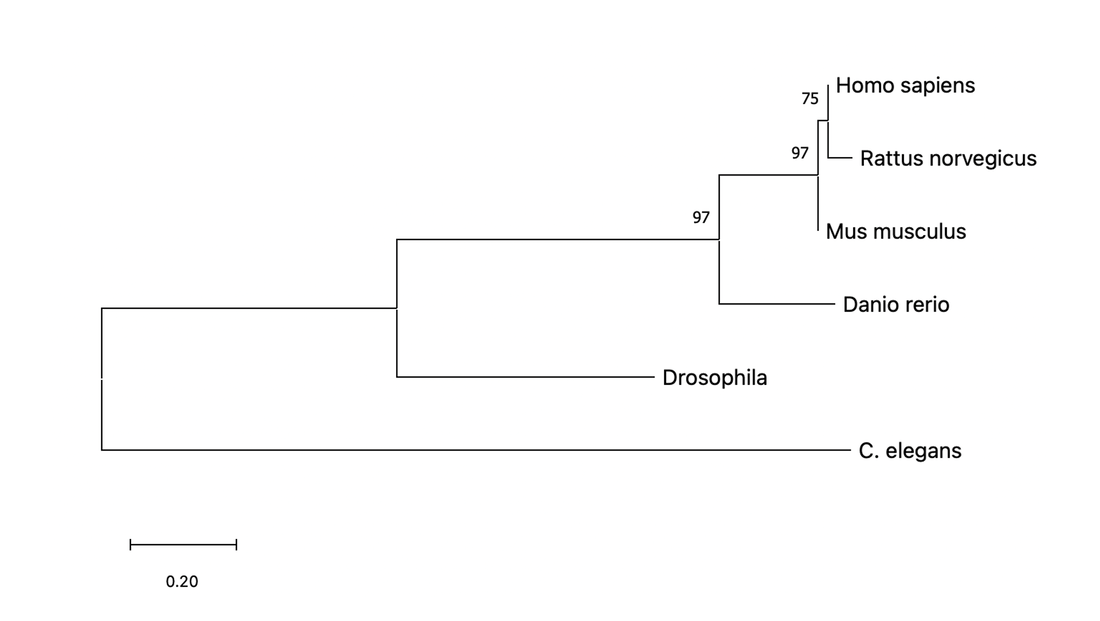
This phylogeny tree constructed in Mega using FASTA sequences shows that the rat and mouse are the closest relatives to humans in regards to SCN1A. Zebrafish are also a close relative.
About this website
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison
Joely Swanson, [email protected]
Last updated April 9th, 2024
Genetics 564 website
Joely Swanson, [email protected]
Last updated April 9th, 2024
Genetics 564 website